| |
SACSegmentationFromNormals planar segmentation
Description: SACSegmentationFromNormals planar segmentationTutorial Level: BEGINNER
Contents
The following launch file starts a nodelet manager together with a VoxelGrid PCL filter nodelet. The input PointCloud2 topic is set to /camera/depth/points.
The default filtering values are set to filter data on the z-axis between 0.01 and 1.5 meters, and downsample the data with a leaf size of 0.01 meters.
The segmentation is performed by first estimating surface normals at each point for a support of 0.015 meters, and then using a RANSAC-based estimator for fitting planes with a threshold of 0.1 meters, and a normal deviation of ~5 degrees (0.09 radians).
1 <launch>
2 <node pkg="nodelet" type="nodelet" name="pcl_manager" args="manager" output="screen" />
3
4 <!-- Run a VoxelGrid filter to clean NaNs and downsample the data -->
5 <node pkg="nodelet" type="nodelet" name="voxel_grid" args="load pcl/VoxelGrid pcl_manager" output="screen">
6 <remap from="~input" to="/camera/depth/points" />
7 <rosparam>
8 filter_field_name: z
9 filter_limit_min: 0.01
10 filter_limit_max: 1.5
11 filter_limit_negative: False
12 leaf_size: 0.01
13 </rosparam>
14 </node>
15
16 <!-- Estimate point normals -->
17 <node pkg="nodelet" type="nodelet" name="normal_estimation" args="load pcl/NormalEstimation pcl_manager" output="screen">
18 <remap from="~input" to="/voxel_grid/output" />
19 <rosparam>
20 # -[ Mandatory parameters
21 k_search: 0
22 radius_search: 0.015
23 # Set the spatial locator. Possible values are: 0 (ANN), 1 (FLANN), 2 (organized)
24 spatial_locator: 0
25 </rosparam>
26 </node>
27
28 <!-- Segment the table plane -->
29 <node pkg="nodelet" type="nodelet" name="planar_segmentation" args="load pcl/SACSegmentationFromNormals pcl_manager" output="screen">
30 <remap from="~input" to="/voxel_grid/output" />
31 <remap from="~normals" to="/normal_estimation/output" />
32 <rosparam>
33 # -[ Mandatory parameters
34 # model_type:
35 # 0: SACMODEL_PLANE
36 # 1: SACMODEL_LINE
37 # 2: SACMODEL_CIRCLE2D
38 # 3: SACMODEL_CIRCLE3D
39 # 4: SACMODEL_SPHERE
40 # 5: SACMODEL_CYLINDER
41 # 6: SACMODEL_CONE
42 # 7: SACMODEL_TORUS
43 # 8: SACMODEL_PARALLEL_LINE
44 # 9: SACMODEL_PERPENDICULAR_PLANE
45 # 10: SACMODEL_PARALLEL_LINES
46 # 11: SACMODEL_NORMAL_PLANE
47 # 12: SACMODEL_NORMAL_SPHERE
48 # 13: SACMODEL_REGISTRATION
49 # 14: SACMODEL_REGISTRATION_2D
50 # 15: SACMODEL_PARALLEL_PLANE
51 # 16: SACMODEL_NORMAL_PARALLEL_PLANE
52 # 17: SACMODEL_STICK
53 model_type: 11
54 distance_threshold: 0.1
55 max_iterations: 1000
56 method_type: 0
57 optimize_coefficients: true
58 normal_distance_weight: 0.1
59 eps_angle: 0.09
60 </rosparam>
61 </node>
62
63 <node pkg="nodelet" type="nodelet" name="extract_plane_indices" args="load pcl/ExtractIndices pcl_manager" output="screen">
64 <remap from="~input" to="/voxel_grid/output" />
65 <remap from="~indices" to="/planar_segmentation/inliers" />
66 <rosparam>
67 negative: true
68 </rosparam>
69 </node>
70 </launch>
In the first terminal,
roscore
Open a new shell, and put following command.
roslaunch openni2_launch openni2.launch roslaunch pcl_ros_tutorials SAC.launch
Then, launch rviz.
rviz
Then, you push "Add" button and choose "PointCloud2".

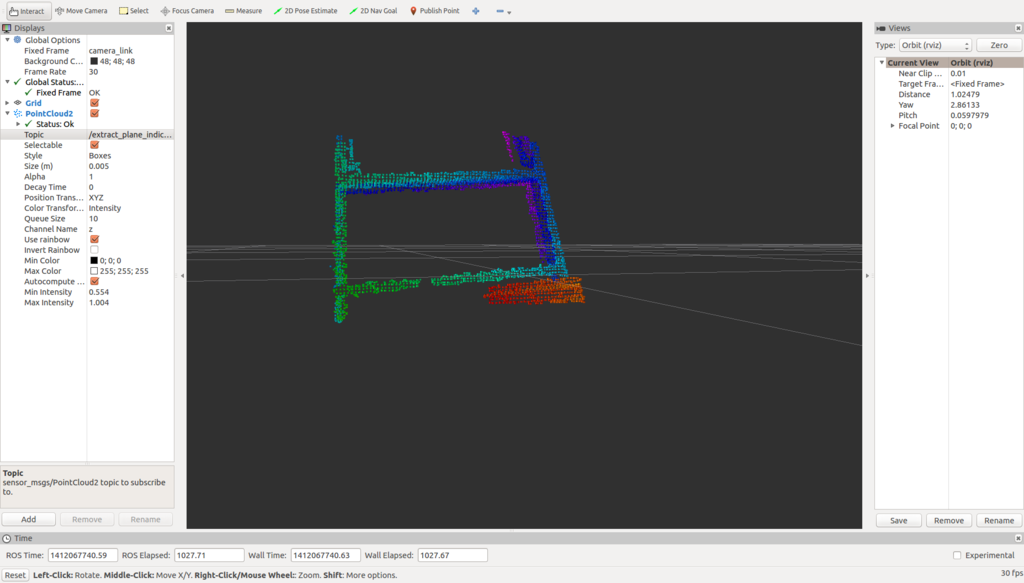
The following image is image extracted plane.